4.10 Models with Covariates#
mode = "svg"
import matplotlib
font = {'family' : 'Dejavu Sans',
'weight' : 'normal',
'size' : 20}
matplotlib.rc('font', **font)
import matplotlib
from matplotlib import pyplot as plt
from graspologic.simulations import sample_edges
import numpy as np
from graphbook_code import heatmap
n = 5
P_earthling = np.full((n,n), 0.3)
nodenames = [
"SI", "L", "H/E",
"T/M", "BS"
]
signal_subnetwork = np.zeros((n, n), dtype=bool)
signal_subnetwork[1:, 0] = True
signal_subnetwork[0, 1:] = True
P_astronaut = np.copy(P_earthling)
# probabilities for signal edges are higher in astronauts than humans
n_different_edges = signal_subnetwork.sum()
P_astronaut[signal_subnetwork] = np.tile(np.linspace(0.35, 0.9, num=n_different_edges//2), 2)
A_earthling = sample_edges(P_earthling)
A_astronaut = sample_edges(P_astronaut)
from graphbook_code import heatmap
import os
fig, axs = plt.subplots(1, 2, figsize=(12, 5), gridspec_kw={"width_ratios": [1, 1.27]})
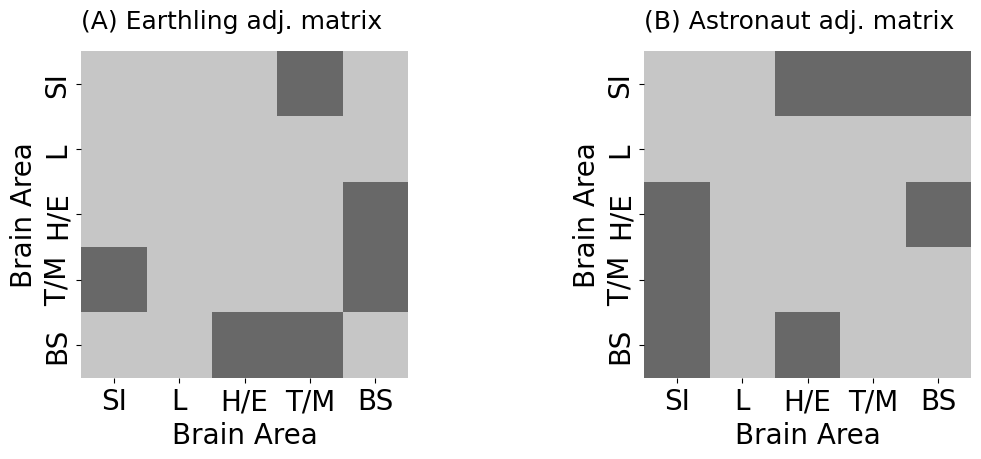
heatmap(A_earthling.astype(int), ax=axs[0], title="(A) Earthling adj. matrix",
xtitle="Brain Area", xticks=[0.5, 1.5, 2.5, 3.5, 4.5], xticklabels=nodenames,
ytitle="Brain Area", yticks=[0.5, 1.5, 2.5, 3.5, 4.5], yticklabels=nodenames,
cbar=False)
heatmap(A_astronaut.astype(int), ax=axs[1], title="(B) Astronaut adj. matrix",
xtitle="Brain Area", xticks=[0.5, 1.5, 2.5, 3.5, 4.5], xticklabels=nodenames,
ytitle="Brain Area", yticks=[0.5, 1.5, 2.5, 3.5, 4.5], yticklabels=nodenames,
cbar=False)
fig.tight_layout()
os.makedirs("Figures", exist_ok=True)
fname = "ssg_samps"
if mode != "png":
os.makedirs(f"Figures/{mode:s}", exist_ok=True)
fig.savefig(f"Figures/{mode:s}/{fname:s}.{mode:s}")
os.makedirs("Figures/png", exist_ok=True)
fig.savefig(f"Figures/png/{fname:s}.png")
fig, axs = plt.subplots(1, 3, figsize=(15, 5), gridspec_kw={"width_ratios": [1, 1.27, 1.27]})
# plot probability matrices and their differences on the same scale
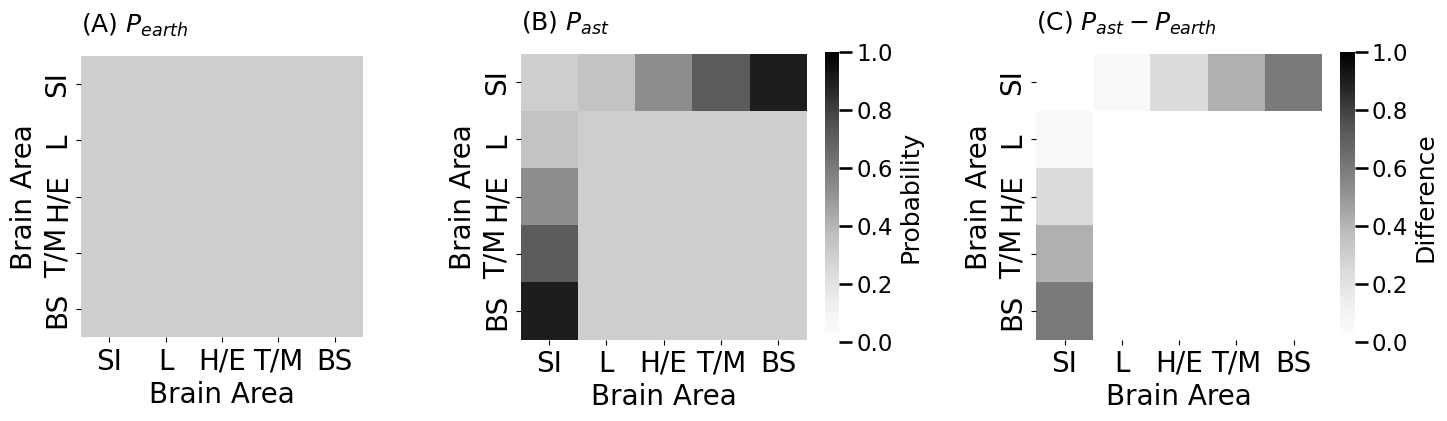
heatmap(P_earthling, vmin=0, vmax=1, title="(A) $P_{earth}$",
xtitle="Brain Area", xticks=[0.5, 1.5, 2.5, 3.5, 4.5], xticklabels=nodenames,
ytitle="Brain Area", yticks=[0.5, 1.5, 2.5, 3.5, 4.5], yticklabels=nodenames,
cbar=False, ax=axs[0])
heatmap(P_astronaut, vmin=0, vmax=1, title="(B) $P_{ast}$",
xtitle="Brain Area", xticks=[0.5, 1.5, 2.5, 3.5, 4.5], xticklabels=nodenames,
ytitle="Brain Area", yticks=[0.5, 1.5, 2.5, 3.5, 4.5], yticklabels=nodenames,
legend_title="Probability", ax=axs[1])
heatmap(P_astronaut - P_earthling, vmin=0, vmax=1, title="(C) $P_{ast} - P_{earth}$",
xtitle="Brain Area", xticks=[0.5, 1.5, 2.5, 3.5, 4.5], xticklabels=nodenames,
ytitle="Brain Area", yticks=[0.5, 1.5, 2.5, 3.5, 4.5], yticklabels=nodenames,
legend_title="Difference", ax=axs[2])
fig.tight_layout()
fname = "ssg_pmtxs"
if mode != "png":
fig.savefig(f"Figures/{mode:s}/{fname:s}.{mode:s}")
fig.savefig(f"Figures/png/{fname:s}.png")
fig, axs = plt.subplots(1, 2, figsize=(12, 5), gridspec_kw = {"width_ratios": [1, 1]})
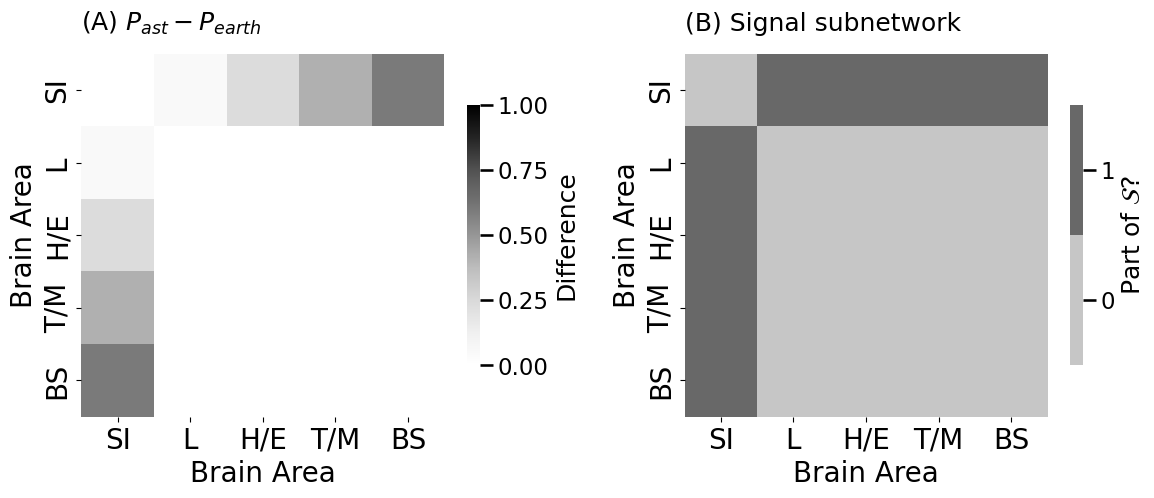
heatmap(P_astronaut - P_earthling, vmin=0, vmax=1, title="(A) $P_{ast} - P_{earth}$",
xtitle="Brain Area", xticks=[0.5, 1.5, 2.5, 3.5, 4.5], xticklabels=nodenames,
ytitle="Brain Area", yticks=[0.5, 1.5, 2.5, 3.5, 4.5], yticklabels=nodenames,
legend_title="Difference", ax=axs[0])
heatmap(signal_subnetwork.astype(int), title="(B) Signal subnetwork",
xtitle="Brain Area", xticks=[0.5, 1.5, 2.5, 3.5, 4.5], xticklabels=nodenames,
ytitle="Brain Area", yticks=[0.5, 1.5, 2.5, 3.5, 4.5], yticklabels=nodenames,
legend_title="Part of $\\mathcal{S}$?", ax=axs[1])
fig.tight_layout()
fname = "ssg_ssn"
if mode != "png":
fig.savefig(f"Figures/{mode:s}/{fname:s}.{mode:s}")
fig.savefig(f"Figures/png/{fname:s}.png")
# sample the classes of each sample
M = 200 # the number of training and testing samples
pi_astronaut = 0.45
pi_earthling = 0.55
yvec = np.random.choice(2, p=[pi_earthling, pi_astronaut], size=M)
# sample network realizations given the class of each sample
Ps = [P_earthling, P_astronaut]
np.random.seed(0)
As = np.stack([sample_edges(Ps[y]) for y in yvec], axis=2)
