C.1 Unsupervised machine learning#
mode = "svg"
import matplotlib
font = {'family' : 'Dejavu Sans',
'weight' : 'normal',
'size' : 20}
matplotlib.rc('font', **font)
import matplotlib
from matplotlib import pyplot as plt
from graspologic.simulations import sbm
from graspologic.embed import AdjacencySpectralEmbed as ASE
import numpy as np
ns = [50, 40, 30]
B = [[0.6, 0.2, 0.2],
[0.2, 0.6, 0.2],
[0.2, 0.2, 0.6]]
np.random.seed(1234)
A = sbm(n=ns, p = B)
# the true community labels
z = [0 for i in range(0,ns[0])] + [1 for i in range(0, ns[1])] + [2 for i in range(0, ns[2])]
Xhat = ASE(n_components=3).fit(A).latent_left_
from pandas import DataFrame
import seaborn as sns
import matplotlib.pyplot as plt
data = DataFrame({"Dimension 2" : Xhat[:,1], "Dimension 3" : Xhat[:,2]})
palette = {"0" : "blue", "1": "green", "2": "red"}
fig, ax = plt.subplots(1, 1, figsize=(6, 4))
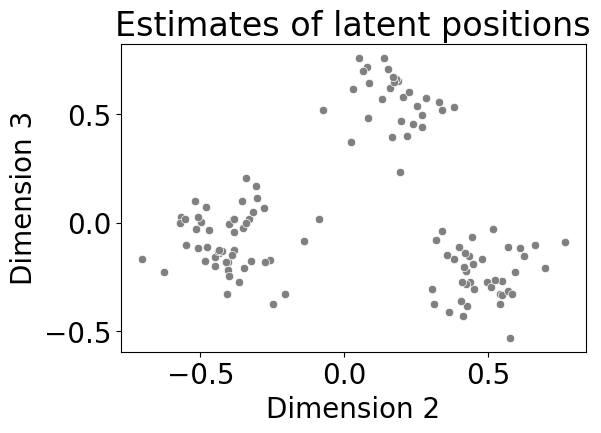
sns.scatterplot(data=data, x="Dimension 2", y="Dimension 3", color="gray", ax=ax)
ax.set_title("Estimates of latent positions");
centers = np.array([[.5, .5], [-0.05, 0.05], [-0.05, -0.05]])
datcenters = DataFrame({"Dimension 2": centers[:,0], "Dimension 3": centers[:,1], "Cluster": ["0", "1","2"]})
from scipy.spatial import distance_matrix
distances = distance_matrix(Xhat[:,1:3], centers)
assignment = np.argmin(distances, axis=1)
data["Closest Center"] = assignment.astype(str)
centers = np.array([np.mean(Xhat[assignment == k,1:3], axis=0) for k in range(0, 3)])
datcenters = DataFrame({"Dimension 2": centers[:,0], "Dimension 3": centers[:,1], "Cluster": ["0", "1","2"]})
distances = distance_matrix(Xhat[:,1:3], centers)
assignment = np.argmin(distances, axis=1)
centers_new = np.array([np.mean(Xhat[assignment == k,1:3], axis=0) for k in range(0, 3)])
data["Closest Center"] = assignment.astype(str)
import os
fig, axs = plt.subplots(1, 3, figsize=(20, 6))
color_kwarg = [{"color": "gray"}, {"hue": "Closest Center"}, {"hue": "Closest Center"}]
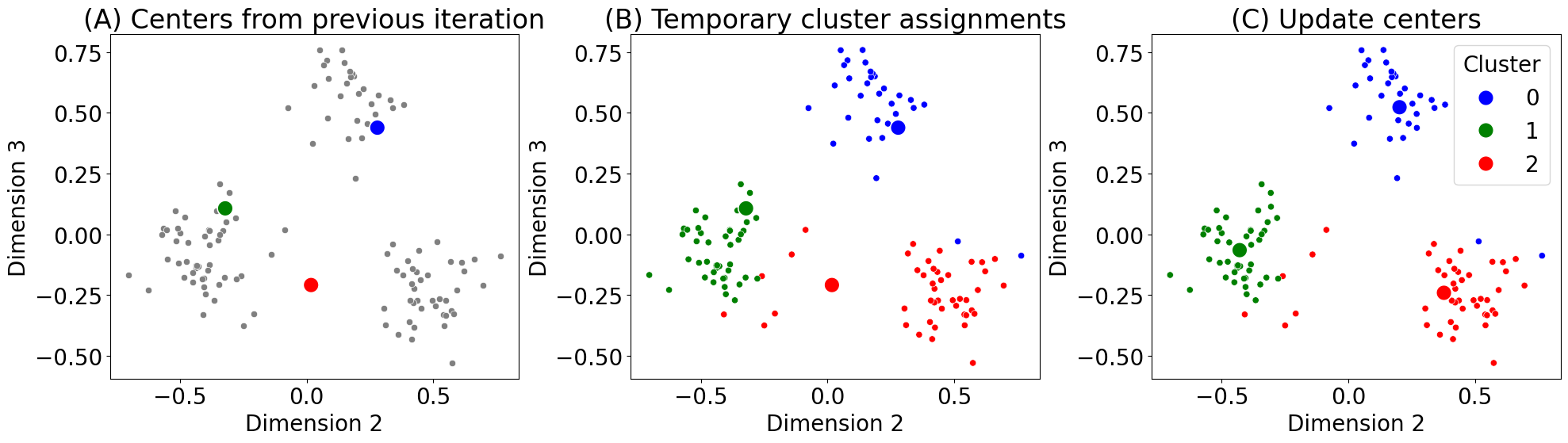
cdat = [centers, centers, centers_new]
titles = ["(A) Centers from previous iteration", "(B) Temporary cluster assignments", "(C) Update centers"]
for i, ax in enumerate(axs.flat):
sns.scatterplot(data=data, x="Dimension 2", y="Dimension 3", ax=ax, **color_kwarg[i],
palette=palette, legend=False)
datcenters = DataFrame({"Dimension 2": cdat[i][:,0], "Dimension 3": cdat[i][:,1], "Cluster": ["0", "1","2"]})
sns.scatterplot(data=datcenters, x="Dimension 2", y="Dimension 3", hue="Cluster",
palette=palette, ax=ax, s=200)
ax.set_title(titles[i])
if i != 2:
ax.get_legend().remove()
fig.tight_layout()
os.makedirs("Figures", exist_ok=True)
fname = "kmeans_process"
if mode != "png":
os.makedirs(f"Figures/{mode:s}", exist_ok=True)
fig.savefig(f"Figures/{mode:s}/{fname:s}.{mode:s}")
os.makedirs("Figures/png", exist_ok=True)
fig.savefig(f"Figures/png/{fname:s}.png")
/tmp/ipykernel_2112/2519076243.py:9: UserWarning: Ignoring `palette` because no `hue` variable has been assigned.
sns.scatterplot(data=data, x="Dimension 2", y="Dimension 3", ax=ax, **color_kwarg[i],
from sklearn.cluster import KMeans
labels_kmeans = KMeans(n_clusters = 3, random_state=1234).fit_predict(Xhat)
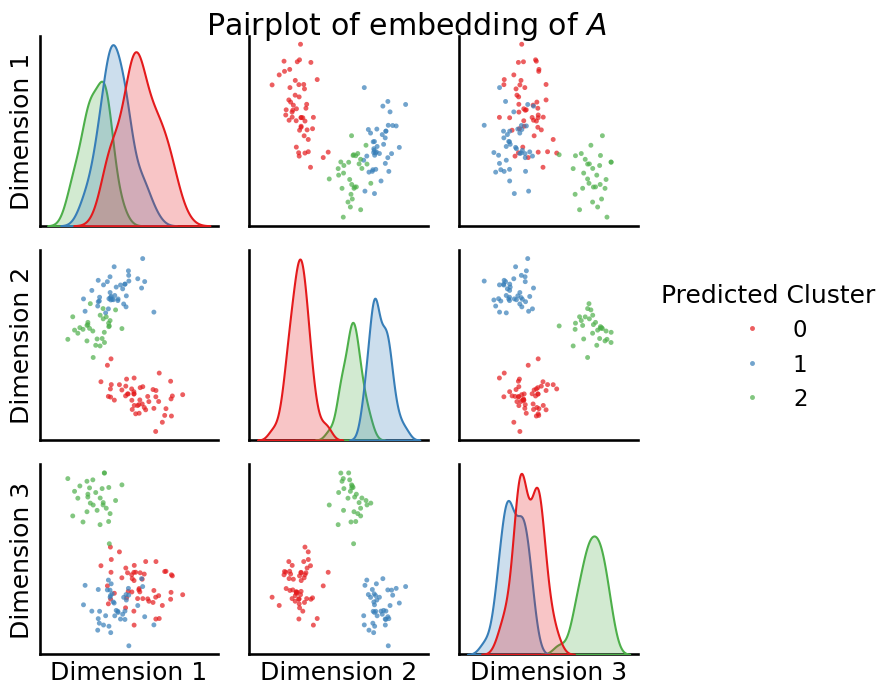
from graspologic.plot import pairplot
fig = pairplot(Xhat, labels=labels_kmeans, title="Pairplot of embedding of $A$", legend_name="Predicted Cluster")
fname = "kmeans_out"
if mode != "png":
fig.savefig(f"Figures/{mode:s}/{fname:s}.{mode:s}")
fig.savefig(f"Figures/png/{fname:s}.png")
from sklearn.metrics import confusion_matrix
# compute the confusion matrix between the true labels z
# and the predicted labels labels_kmeans
cf_matrix = confusion_matrix(z, labels_kmeans)
cfm_norm = cf_matrix/cf_matrix.sum(axis=1)[:,None]
from graphbook_code import cmaps
fig, axs = plt.subplots(1,2, figsize=(12,5))
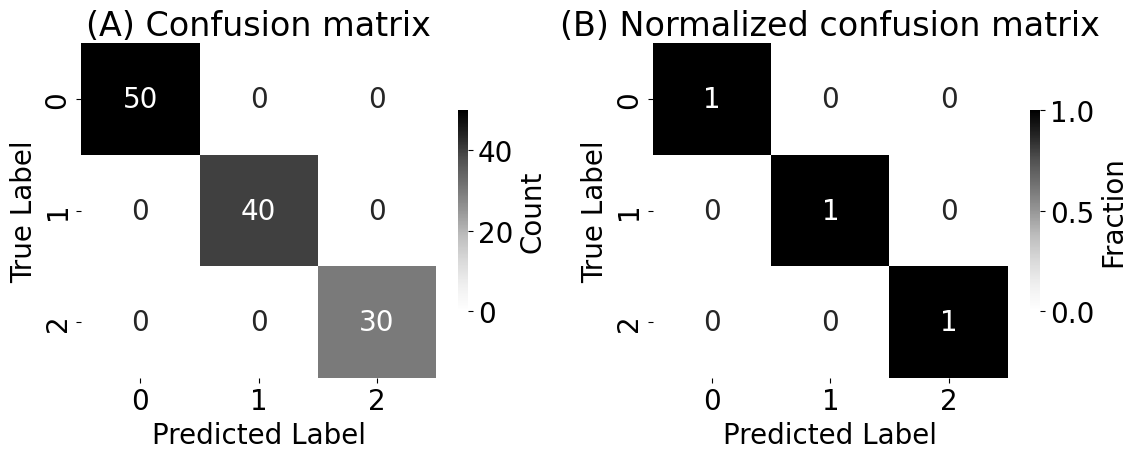
sns.heatmap(cf_matrix, cmap=cmaps["sequential"], ax=axs[0], annot=True, cbar_kws={"label": "Count", "shrink": 0.6})
axs[0].set_title("(A) Confusion matrix")
axs[0].set_ylabel("True Label")
axs[0].set_xlabel("Predicted Label")
sns.heatmap(cfm_norm, cmap=cmaps["sequential"], ax=axs[1], annot=True, cbar_kws={"label": "Fraction", "shrink": 0.6})
axs[1].set_title("(B) Normalized confusion matrix")
axs[1].set_ylabel("True Label")
axs[1].set_xlabel("Predicted Label")
fig.tight_layout()
fname = "kmeans_cfmtx"
if mode != "png":
fig.savefig(f"Figures/{mode:s}/{fname:s}.{mode:s}")
fig.savefig(f"Figures/png/{fname:s}.png")
from sklearn.metrics import adjusted_rand_score
ari_kmeans = adjusted_rand_score(z, labels_kmeans)
print("ARI(predicted communities, true communities) = {}".format(ari_kmeans))
ARI(predicted communities, true communities) = 1.0
